|
A maximum likelihood approach was used to estimate the parameters of different prediction equations. Different k-fold (k = 2–10, 15, 20) cross-validation scenarios (50 replicates, random assignment) were performed using a genomic BLUP approach. Two data sets of 5′698 Holstein Friesian bulls genotyped with 50 K SNPs and 1′332 Brown Swiss bulls genotyped with 50 K SNPs and imputed to ∼600 K SNPs were available. 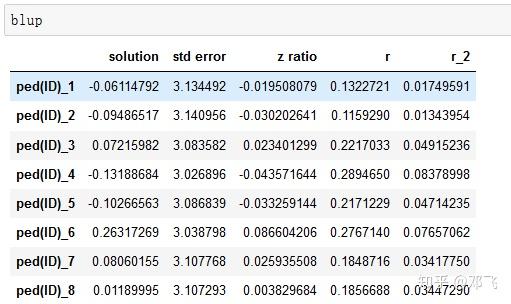
The aim of our study was to find a general deterministic equation for the average accuracy of genomic breeding values that also accounts for marker density and can be fitted empirically. 
Deterministic equations have been suggested to predict the accuracy of genomic breeding values in a given design which are based on training set size, reliability of phenotypes, and the number of independent chromosome segments ( ). Prediction of genomic breeding values is of major practical relevance in dairy cattle breeding.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
